An Integrated Approach to Characterizing Conformational Changes of Large Proteins
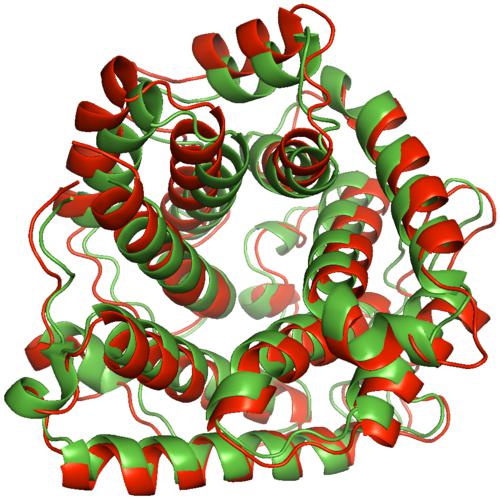
One of the major hurdles in the understanding of physiological processes and the design of new pharmaceutical drugs has been the understanding of the three-dimensional structure and flexibility of molecules, which hold key insight into their function and interactions. In many biological pathways conformational flexibility is known to play an essential role in driving functional mechanisms and has been the topic of both computational and experimental investigations. Modern experimental methods such as Hydrogen/Deuterium Exchange coupled with Mass Spectroscopy (HDX-MS) can probe the flexibility of large proteins, but the findings are mapped on the available (static) crystal structures, thereby losing a considerable part of the dynamic information. This project will investigate the extent to which computational methods can help discover and visualize conformational changes of large proteins that are in agreement with experiment. Because of the specific proteins selected for study in the context of this project, the proposed work has a great societal impact: it helps advance the understanding of processes that are involved in a variety of immune, inflammatory, neurodegenerative, ischemic, and age-related diseases. This project will train undergraduate, graduate, and postdoctoral students at Rice and the University of Pennsylvania on interdisciplinary topics. It will also provide training opportunities to women students. All work will be disseminated not only through technical presentations at national meetings, publications in peer reviewed journals and book chapters, but also through visits and talks with researchers in academia and the biotechnology industry, as well as with open source development and distribution of the algorithms developed for widespread use. A central part of this project concerns the development of a computational framework that samples the conformational space of proteins. It relies on a new representation of the degrees of freedom of proteins and protein complexes and on the definition of sampling primitives at various resolutions. The framework draws from geometry-inspired sampling techniques that efficiently explore large search spaces while keeping track of the extent that the search space is covered. In parallel, HDX-MS experiments will be carried out and used as probes for conformational variability. The experimental results will be correlated with the results of the computational framework to select the conformations that most closely match experiments. A distinguishing feature of this work is the connection with experiment and the ways in which each can feed the other. This project lies at the intersection of algorithms, computational geometry, dimensionality reduction, data mining, and visualization methods and connects with modern experimental methods which are actively under development and hold great promise for the understanding of protein structure and function.
This work has been supported by grant NSF AF 1423304.
Related Publications
- D. Devaurs, D. A. Antunes, and L. E. Kavraki, “Computational analysis of complement inhibitor compstatin using molecular dynamics,” Journal of Molecular Modeling, vol. 26, no. 231, Aug. 2020.
Details - E. Hruska, J. R. Abella, F. Nüske, L. E. Kavraki, and C. Clementi, “Quantitative comparison of adaptive sampling methods for protein dynamics,” The Journal of Chemical Physics, vol. 149, no. 24, p. 244119, Dec. 2018. PMID: 30599712
Details - D. Devaurs, M. Papanastasiou, D. A. Antunes, J. R. Abella, M. Moll, D. Ricklin, J. D. Lambris, and L. E. Kavraki, “Native state of complement protein C3d analysed via hydrogen exchange and
conformational sampling,” International Journal of Computational Biology and Drug Design, vol. 11, no. 1/2, pp. 90–113, 2018. PMID: 30700993, PMCID: PMC6349257
Details - D. Devaurs, D. A. Antunes, and L. E. Kavraki, “Revealing Unknown Protein Structures Using Computational Conformational Sampling Guided
by Experimental Hydrogen-Exchange Data,” International Journal of Molecular Sciences, vol. 19, no. 11, p. 3406, 2018. PMID: 30384411, PMCID: PMC6280153
Details - J. R. Abella, M. Moll, and L. E. Kavraki, “Maintaining and Enhancing Diversity of Sampled Protein Conformations in
Robotics-Inspired Methods,” Journal of Computational Biology, vol. 25, no. 1, pp. 3–20, 2018. PMID: 29035572, PMCID: PMC5756939
Details - A. Novinskaya, D. Devaurs, M. Moll, and L. E. Kavraki, “Defining low-dimensional projections to guide protein conformational sampling,” Journal of Computational Biology, vol. 24, no. 1, pp. 79–89, 2017. PMID: 27892695
Details - D. Devaurs, D. A. Antunes, M. Papanastasiou, M. Moll, D. Ricklin, J. D. Lambris, and L. E. Kavraki, “Coarse-grained conformational sampling of protein structure improves the fit to
experimental hydrogen-exchange data,” Frontiers in Molecular Biosciences, vol. 4, no. 13, 2017. PMCID: PMC5344923, PMID: 28344973
Details - M. Moll, P. W. Finn, and L. E. Kavraki, “Structure-Guided Selection of Specificity Determining Positions in the Human Kinome,” BMC Genomics, vol. 17 (Suppl. 4), p. 431, 2016. PMID: 27556159, PMCID: PMC5001202
Details - A. Novinskaya, D. Devaurs, M. Moll, and L. E. Kavraki, “Improving protein conformational sampling by using guiding projections,” in Proceedings of the IEEE International Conference on Bioinformatics and Biomedicine
Workshops (BIBMW), 2015, pp. 1272–1279.
Details - M. Moll, P. W. Finn, and L. E. Kavraki, “Structure-Guided Selection of Specificity Determining Positions in the Human Kinome,” in Proceedings of the IEEE International Conference on Bioinformatics and Biomedicine
(BIBM), 2015, pp. 21–28.
Details - D. A. Antunes, D. Devaurs, and L. E. Kavraki, “Understanding the challenges of protein flexibility in drug design,” Expert Opinion on Drug Discovery, vol. 10, no. 12, pp. 1301–1313, 2015. PMID: 26414598
Details